In January 2015 I was invited by Ms. Henriette van der Zwan to become associated study leader of her PDH study to sequence the genome of genus Agapornis.
The first species on the list was Agapornis roseicollis and in 2016 we collected blood samples of these birds. The genome of Agapornis roseicollis has been sequenced in 2017. (DNA sequencing is the process of determining the precise order of nucleotides within a DNA molecule).
Our first article on the lovebird genome “Draft De Novo Genome Sequence of Agapornis roseicollis for Application in Avian Breeding” by Henriëtte van der Zwan, Francois van der Westhuizen, Carina Visser & Rencia van der Sluis” was published in October 2017. Please feel free to read it here:
Currently, we are busy developing the parentage verification test for all lovebird species. This will allow breeders and buyers to verify the parents of a bird before it is sold.
We also hope to develop a test to determine if a young is a split for a specific color mutation, even before it is sold. This will, however, be in a later stage of the study and this will be the hardest part since it has never been done before and it will probably take a few years.
This will cost a lot of money. Therefore we need to get some extra money and we will soon start fund raising. We are also in urgent need of DNA Samples of all Agapornis species.
If you or your society might be able to help, you can find all info on this link.
Agapornis Genome Study by Henriëtte van der Zwan
Introduction
Agapornis or Lovebird breeding is a worldwide industry and yet very little molecular genetics research has been done on these species. The North-West University in Potchefstroom, South Africa has started a study to sequence the genome of the lovebird that will allow the identification of markers useful to verify parentage and to identify the mutations causing colour variations.
ABC of DNA
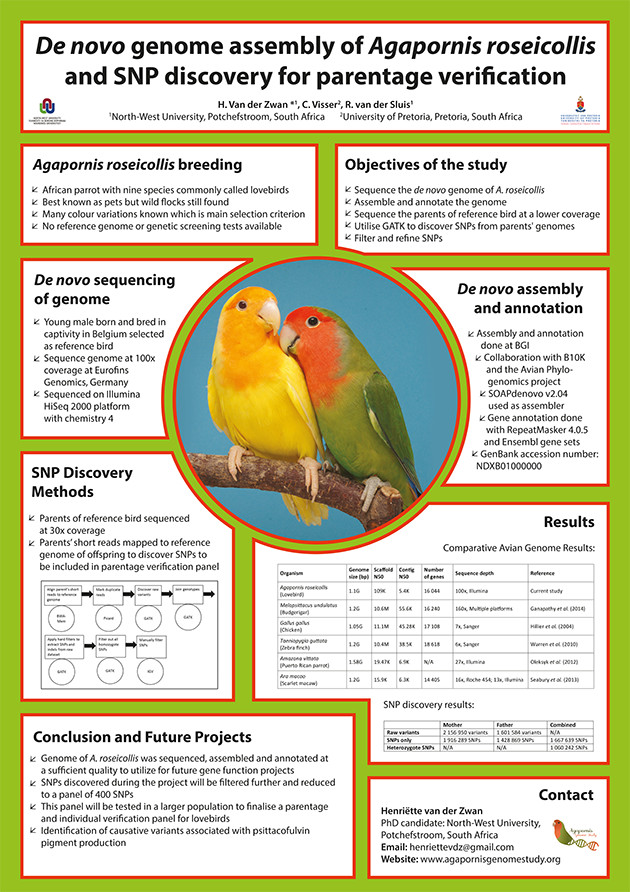
DNA is the molecule in each and every living organism that carries the hereditary traits of the organism. The DNA structure of humans, birds, animals, plants, viruses and bacteria are all the same. DNA is densely packaged into chromosomes and each species has a different number of chromosomes. If we could pull the chromosome apart we will find a double helix structure that is the DNA. Imagine the DNA molecule as a spiral staircase. Each step of the staircase is a base pair. There are only four bases (A, T, G and C) and these always bind as A-T and C-G to form the base pair.
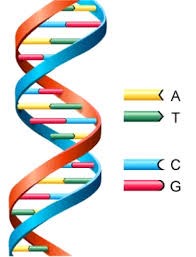
Figure 1: DNA molecule showing bases A, T, C and G as well as base pairs
The DNA molecule is made up of areas of genes and areas of “junk DNA”. The largest part of the DNA is junk DNA and scientists are still trying to understand the full role of these area.
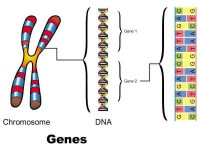
Figure 2: Chromosome, DNA structure with two genes and the base sequence of Gene 2
The sequence in which the bases are arranged are fixed for a specific species. However there are areas where the bases differ and this causes genetic diversity. This is called a mutation. A mutation can have either a positive or negative effect. A positive effect includes a colour variant where a negative effect can lead to an inherited disease or abnormality. This mutation can simply be a substitution of one nucleotide for another (e.g. an A for a C) or it can be a deletion or addition of a nucleotide into a gene.
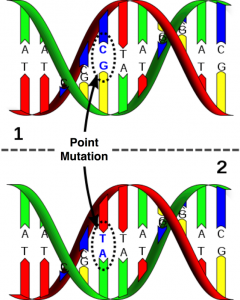
Figure 3: DNA molecule with a mutation
Genes and markers
The use of the terms “genes” and “markers” is often confusing. The easiest illustration of the difference between a gene and a marker is to imagine the DNA as a long stretch of road. On your travels you will pass through areas where there are towns and areas where there are open fields. Imagine the towns as being “genes” and the open areas as “junk DNA”. The towns (or genes) have a specific sequence – there is the supermarket, then the hairdresser, then the liquor store etc. If the hairdresser and the liquor store suddenly trade places it will cause mayhem and the clients won’t know what to buy (thus the same as what happens with a mutation). The open areas (or junk DNA) have no specific function. However there may be a big rock or hill 1.2 km outside of the town. This acts as a marker – it is a fixed object that always look the same and always at the same spot. The rock doesn’t have a specific function, but it acts as a marker of location.
A gene is a segment of DNA with a specific function (like the town) and it needs to be in a specific sequence to be functional. Its function is to produce proteins that are again used in different physiological processes. For example the genes involved in feather colour pigmentation has the function of sending a message to produce pigment for the feathers. If the sequence in the gene is changed then the message gets messed up and the pigment isn’t expressed in the right way, leading to colour variations.
Markers are mostly found in the junk DNA areas. There are different types of markers but the two most commonly used are Single Nucleotide Polymorphisms (SNPs) and microsatellites (STRs). These are differences between individuals but it doesn’t have a specific influence on a gene or any physiological process. We make use of SNPs as genetic markers because these changes (polymorphisms) of one (single) base (nucleotide) is inherited from parent to offspring allowing us to identify which individual has sired which offspring. For a parentage verification test the parents and the offspring need to be tested at the panel of SNPs and the data of the three animals are compared. If the male is the sire the chick had to inherit one copy from the male and if the female is the dam it had to inherit one copy from her.
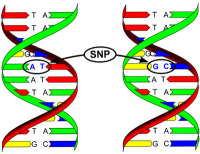
Figure 4: A SNP marker is similar to a mutation, but the change in the DNA has no influence on any physiological process
What the study will be about
During this study we will sequence the full genome of one Agapornis roseicollis offspring. What this mean is we will determine the sequence of all the base pairs of the DNA. This will give us the blueprint of what a roseicollis genome looks like. This has never been done anywhere in the world for any lovebird species. Thereafter we want to re-sequence the parents of this offspring. They will act as the controls to verify that the offspring’s genome was correctly assembled.
Parentage verification panel
After the three bird’s genomes are assembled, we will identify SNP markers that can be used as a panel to verify parentage and also identity. This will allow the breeder to sell a chick with a DNA profile and also the guarantee that a specific parent pair is the biological parents of the chick. We want to offer this as a test to all lovebird species, not only roseicollis.
Colour mutations
From the data generated in the genome assembly we can identify the mutations in the colour genes causing different colour variations. We want to assemble a test that allows breeders to test a chick and know for certain that it is a split (or heterozygote) for a specific recessive colour mutation. The parents won’t have to be tested (only the chick) and it will give you a reliable genetic profile of the bird’s colour inheritance. We hope to offer this test for as many different colour mutations over all the species as possible, however we cannot guarantee that all mutations will be identified.
Future plans
This phase of the study will only include roseicollis individuals but we hope to test the other Agapornis species too to verify the colour tests as well as parentage markers in all the species.
How you can help us:
We are in need of samples! We need to identify a parent pair and offspring that we can use in this study. We want to keep the birds alive and only use blood samples, hopefully only drawing blood once. The birds will remain with the owner and we will only use the blood samples taken by a chosen veterinarian. If the birds should die during the study we would like to preserve the carcass should we need more DNA at a later stage.
If you think you can help us please contact us by email
More info: www.agapornisgenomestudy.org

